Multiscale modelling, integrating electrical and biochemical
A model can be considered multiscale when one or both these conditions are true: (i) the object of the modelling spawns different orders of magnitude in at least time or space e.g.: from microseconds (μs) to minutes (min), or from nanometers (nm) to millimeters (mm); (ii) parts of the model run with different timescales and/or with different spatial resolutions, influencing each other through a systematic exchange of variables (Weinan and Engquist, 2003).
The result of the simulation is the summa of the different approaches, combined together, and cannot be achieved using one part of the model only, or simulating them in a disconnected way. All branches of Biology are inherently multiscale. Neuroscience as well has also to address multiscale problems, where for example, an excitatory release of glutamate can trigger an increase in the Calcium concentration, which can then act as second messenger, stimulating complex biochemical cascades with timescales spanning from minutes to hours or days. Nonetheless, given the wide spectrum of the processes involved, communities have focused on one particular domain, the electrical aspect or the biochemical one.
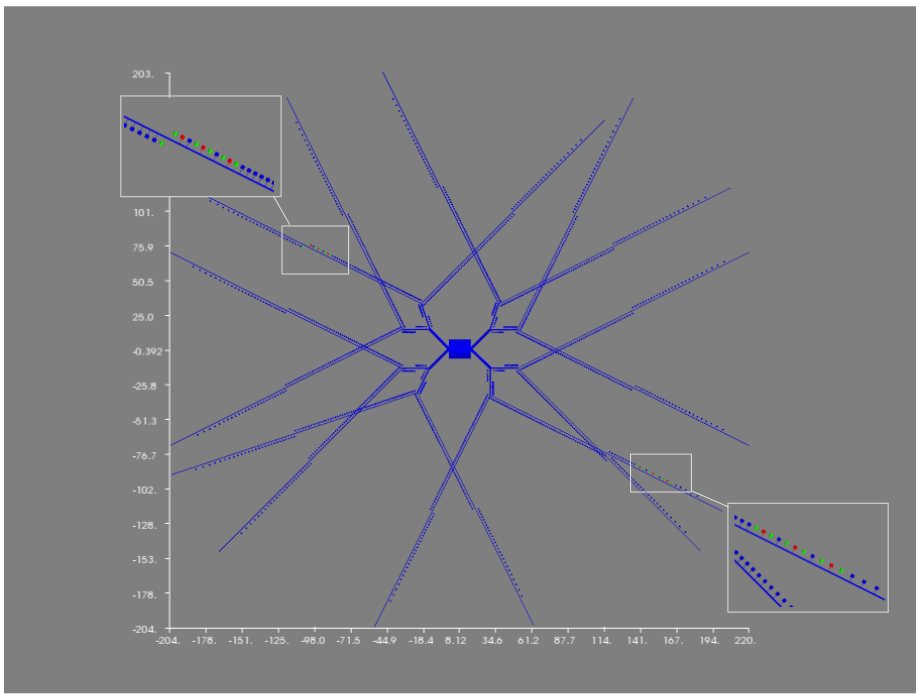
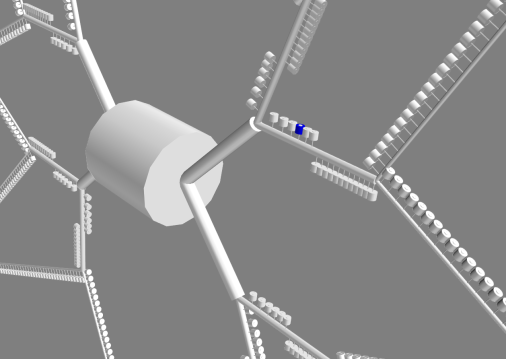
In my Ph.D. I have developed an event-driven algorithm which allows to integrate different simulators, optimizing the number of syncs between the two and the time of the syncing. At the same time I’ve also developed a multiscale model of a Medium Spiny Neuron of the Striatum, with 1504 spine modelled explicitely where the plasticity, the strenght of a synapse, is computed as the results of the integration between the electrical and the biochemical.
The paper about the project can be found on Plos One, the code is on Github, and the documentation is available at Timescale documentation.